PEAK 15 (8.191 ppm) DETAILS
GENERAL INFORMATION
PEAK DETECTION

WINDOW ALIGNMENT

INTENSITY DISTRIBUTION
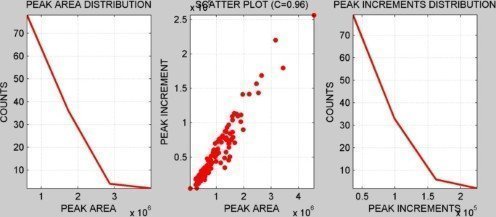
CORRELATION ANALYSIS

| CORRCOEF | ID_PEAK | INFO | PPM | ID_WINDOW | QC_SC | QC_IvsA | QC_IP | G_95 | G_90 | G_85 | G_80 | INFO | |||
| 0.79 | p22 | + | 7.432 | 94 (7.462-7.384) | 0.99 | 0.97 | 0.25 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.8 | p23 | + | 7.42 | 94 (7.462-7.384) | 0.98 | 0.95 | 0.15 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.8 | p25 | + | 7.34 | 96 (7.384-7.306) | 0.98 | 0.93 | 0.3 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.79 | p26 | + | 7.327 | 96 (7.384-7.306) | 0.98 | 0.9 | 0.15 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.83 | p30 | + | 7.205 | 100 (7.228-7.15) | 0.98 | 0.93 | 0.35 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.82 | p31 | + | 7.191 | 100 (7.228-7.15) | 0.98 | 0.93 | 0.4 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.78 | p32 | + | 7.06 | 103 (7.11-7.032) | 0.91 | 0.93 | 0.15 | G95_30 (1) | G90_23 (1) | G85_1 (69) | G80_1 (106) | + | |||
| 0.83 | p34 | + | 6.909 | 107 (6.954-6.876) | 0.98 | 0.95 | 0.3 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.82 | p35 | + | 6.895 | 107 (6.954-6.876) | 0.98 | 0.93 | 0.3 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.79 | p89 | + | 4.256 | 175 (4.294-4.216) | 0.59 | 0.5 | 0.1 | G95_39 (1) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.72 | p240 | + | 3.349 | 198 (3.395-3.317) | 0.88 | 0.76 | 0.1 | G95_80 (0) | G90_57 (0) | G85_1 (69) | G80_1 (106) | + | |||
| 0.71 | p270 | + | 3.134 | 204 (3.16-3.082) | 0.49 | 0.66 | 0.25 | G95_90 (0) | G90_66 (0) | G85_1 (69) | G80_1 (106) | + | |||
| 0.8 | p273 | + | 3.069 | 206 (3.082-3.004) | 0.75 | 0.74 | 0.25 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.85 | p274 | + | 3.057 | 206 (3.082-3.004) | 0.76 | 0.81 | 0.5 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.83 | p276 | + | 3.029 | 206 (3.082-3.004) | 0.98 | 0.88 | 0.6 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.86 | p277 | + | 3.016 | 206 (3.082-3.004) | 0.96 | 0.81 | 0.55 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.71 | p298 | + | 2.83 | 211 (2.886-2.808) | 0.72 | 0.67 | 0.15 | G95_94 (0) | G90_10 (3) | G85_1 (69) | G80_1 (106) | + | |||
| 0.71 | p303 | + | 2.68 | 216 (2.691-2.613) | 0.86 | 0.86 | 0.2 | G95_19 (2) | G90_10 (3) | G85_1 (69) | G80_1 (106) | + | |||
| 0.79 | p304 | + | 2.648 | 216 (2.691-2.613) | 0.95 | 0.9 | 0.5 | G95_20 (2) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.8 | p305 | + | 2.635 | 216 (2.691-2.613) | 0.94 | 0.83 | 0.3 | G95_20 (2) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.81 | p363 | + | 1.754 | 239 (1.791-1.713) | 0.91 | 0.72 | 0.3 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.83 | p366 | + | 1.742 | 240 (1.752-1.674) | 0.92 | 0.77 | 0.3 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.83 | p367 | + | 1.728 | 240 (1.752-1.674) | 0.95 | 0.8 | 0.4 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.8 | p368 | + | 1.716 | 240 (1.752-1.674) | 0.95 | 0.8 | 0.5 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.8 | p370 | + | 1.705 | 241 (1.713-1.635) | 0.94 | 0.72 | 0.35 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.83 | p384 | + | 1.053 | 257 (1.087-1.009) | 0.96 | 0.83 | 0.7 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.82 | p385 | + | 1.041 | 257 (1.087-1.009) | 0.96 | 0.82 | 0.7 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.82 | p386 | + | 1.022 | 257 (1.087-1.009) | 0.97 | 0.67 | 0.55 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.87 | p389 | + | 1.01 | 258 (1.048-0.97) | 0.96 | 0.69 | 0.5 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.81 | p392 | + | 0.978 | 258 (1.048-0.97) | 0.98 | 0.83 | 0.75 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.83 | p393 | + | 1.0 | 259 (1.009-0.931) | 0.95 | 0.78 | 0.7 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.82 | p394 | + | 0.989 | 259 (1.009-0.931) | 0.93 | 0.74 | 0.7 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.8 | p396 | + | 0.966 | 259 (1.009-0.931) | 0.98 | 0.89 | 0.85 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.81 | p397 | + | 0.956 | 259 (1.009-0.931) | 0.98 | 0.82 | 0.75 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.82 | p398 | + | 0.945 | 259 (1.009-0.931) | 0.96 | 0.65 | 0.55 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + | |||
| 0.73 | p402 | + | 0.932 | 260 (0.97-0.892) | 0.68 | 0.49 | 0.2 | G95_1 (27) | G90_2 (29) | G85_1 (69) | G80_1 (106) | + |
IDENTIFICATION ANALYSIS
| METABOLITE | TYPE | IC:MATCHING | IC:TOTAL | EC:MATCHING | EC:TOTAL | OTHERS | X | Y | Z | SCORE | FIG | |||
| histidine-7.0 (c:1/p:1) | M | 1 | 1 | 1 | 4 | 0 | 0.51 | 0.3 | 1.0 | 0.6 | ||||
| NAD-7.4 (c:6/p:2) | M | 1 | 5 | 1 | 10 | 0 | -0.15 | 0.21 | 1.0 | 0.36 | ||||
| NAD-7.4 (c:6/p:3) | M | 1 | 5 | 1 | 10 | 0 | -0.05 | 0.21 | 1.0 | 0.39 | ||||
| NAD-7.4 (c:6/p:4) | M | 1 | 5 | 1 | 10 | 0 | -0.17 | 0.21 | 1.0 | 0.35 |