PEAK 187 (3.625 ppm) DETAILS
GENERAL INFORMATION
PEAK DETECTION

WINDOW ALIGNMENT

INTENSITY DISTRIBUTION
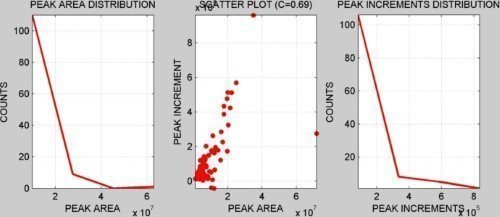
CORRELATION ANALYSIS

| CORRCOEF | ID_PEAK | INFO | PPM | ID_WINDOW | QC_SC | QC_IvsA | QC_IP | G_95 | G_90 | G_85 | G_80 | INFO | |||
| 0.82 | p56 | + | 5.421 | 145 (5.468-5.39) | 0.23 | 0.97 | 0.05 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.96 | p58 | + | 5.404 | 145 (5.468-5.39) | 0.06 | 0.97 | 0.0 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.75 | p60 | + | 5.409 | 146 (5.429-5.351) | 0.2 | 0.88 | 0.0 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.87 | p71 | + | 4.667 | 165 (4.685-4.607) | 0.58 | 0.99 | 0.0 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.88 | p103 | + | 3.995 | 181 (4.06-3.982) | 0.44 | 0.76 | 0.35 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.91 | p106 | + | 3.98 | 182 (4.02-3.943) | 0.59 | 0.88 | 0.6 | G95_12 (2) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.87 | p112 | + | 3.961 | 183 (3.981-3.903) | 0.62 | 0.92 | 0.6 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.81 | p116 | + | 3.924 | 183 (3.981-3.903) | 0.81 | 0.89 | 0.4 | G95_47 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.94 | p128 | + | 3.871 | 185 (3.903-3.825) | 0.26 | 0.92 | 0.1 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.92 | p129 | + | 3.866 | 185 (3.903-3.825) | 0.03 | 0.94 | 0.05 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.89 | p132 | + | 3.845 | 185 (3.903-3.825) | 0.29 | 0.86 | 0.25 | G95_49 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.8 | p135 | + | 3.85 | 186 (3.864-3.786) | 0.91 | 0.96 | 0.7 | G95_51 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.8 | p154 | + | 3.76 | 188 (3.786-3.708) | 0.3 | 0.96 | 0.65 | G95_61 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.88 | p169 | + | 3.683 | 189 (3.747-3.669) | 0.64 | 0.48 | 0.35 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.81 | p170 | + | 3.676 | 189 (3.747-3.669) | 0.73 | 0.59 | 0.55 | G95_65 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.95 | p172 | + | 3.695 | 190 (3.708-3.63) | 0.48 | 0.61 | 0.05 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.77 | p177 | + | 3.66 | 190 (3.708-3.63) | 0.89 | 0.67 | 0.8 | G95_66 (1) | G90_49 (1) | G85_5 (6) | G80_1 (106) | + | |||
| 0.94 | p189 | + | 3.609 | 191 (3.668-3.591) | 0.34 | 0.78 | 0.4 | G95_12 (2) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.91 | p190 | + | 3.6 | 191 (3.668-3.591) | 0.56 | 0.93 | 0.05 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.84 | p192 | + | 3.615 | 192 (3.629-3.551) | 0.76 | 0.74 | 0.45 | G95_69 (1) | G90_51 (0) | G85_2 (39) | G80_1 (106) | + | |||
| 0.86 | p195 | + | 3.589 | 192 (3.629-3.551) | 0.34 | 0.81 | 0.4 | G95_70 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.88 | p249 | + | 3.281 | 200 (3.316-3.238) | 0.7 | 0.81 | 0.45 | G95_4 (8) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + |
IDENTIFICATION ANALYSIS
| METABOLITE | TYPE | IC:MATCHING | IC:TOTAL | EC:MATCHING | EC:TOTAL | OTHERS | X | Y | Z | SCORE | FIG | |||
| glycerol-7.0 (c:2/p:3) | M | 1 | 4 | 1 | 3 | 0 | 0.55 | 0.44 | 1.0 | 0.67 | ||||
| glycerol-7.0 (c:2/p:4) | M | 1 | 4 | 1 | 3 | 0 | 0.36 | 0.44 | 1.0 | 0.6 | ||||
| threonine-7.0 (c:2/p:1) | M | 1 | 2 | 1 | 3 | 0 | 0.51 | 0.23 | 1.0 | 0.58 | ||||
| valine-7.0 (c:1/p:1) | M | 1 | 2 | 1 | 3 | 0 | 0.48 | 0.24 | 1.0 | 0.57 | ||||
| valine-7.0 (c:1/p:2) | M | 1 | 2 | 1 | 3 | 0 | 0.52 | 0.24 | 1.0 | 0.58 |