PEAK 215 (3.481 ppm) DETAILS
GENERAL INFORMATION
PEAK DETECTION

WINDOW ALIGNMENT

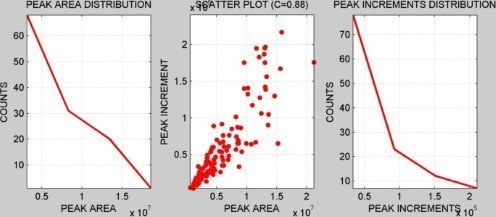
INTENSITY DISTRIBUTION
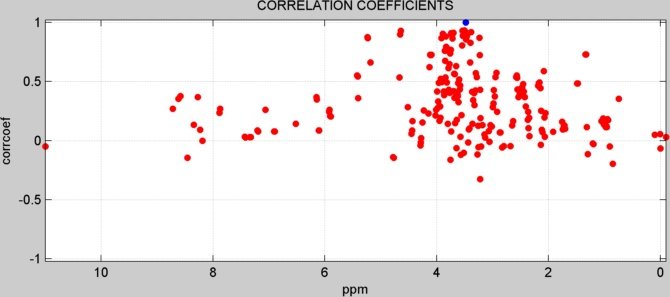
CORRELATION ANALYSIS
| CORRCOEF | ID_PEAK | INFO | PPM | ID_WINDOW | QC_SC | QC_IvsA | QC_IP | G_95 | G_90 | G_85 | G_80 | INFO | |||
| 0.87 | p61 | + | 5.238 | 150 (5.272-5.194) | 0.98 | 0.99 | 0.8 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.87 | p62 | + | 5.231 | 150 (5.272-5.194) | 0.98 | 0.99 | 0.8 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.9 | p72 | + | 4.654 | 165 (4.685-4.607) | 0.99 | 0.99 | 0.9 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.93 | p73 | + | 4.642 | 165 (4.685-4.607) | 0.99 | 0.98 | 0.9 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.73 | p95 | + | 4.118 | 178 (4.177-4.099) | 0.96 | 0.94 | 0.8 | G95_7 (5) | G90_6 (6) | G85_4 (6) | G80_1 (106) | + | |||
| 0.72 | p98 | + | 4.105 | 179 (4.138-4.06) | 0.98 | 0.93 | 0.75 | G95_7 (5) | G90_6 (6) | G85_4 (6) | G80_1 (106) | + | |||
| 0.72 | p99 | + | 4.094 | 179 (4.138-4.06) | 0.97 | 0.83 | 0.5 | G95_7 (5) | G90_6 (6) | G85_4 (6) | G80_1 (106) | + | |||
| 0.87 | p121 | + | 3.913 | 184 (3.942-3.864) | 0.93 | 0.97 | 0.75 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.89 | p126 | + | 3.893 | 185 (3.903-3.825) | 0.97 | 0.96 | 0.8 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.9 | p127 | + | 3.888 | 185 (3.903-3.825) | 0.92 | 0.95 | 0.8 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.88 | p133 | + | 3.836 | 185 (3.903-3.825) | 0.95 | 0.96 | 0.85 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.76 | p134 | + | 3.855 | 186 (3.864-3.786) | 0.9 | 0.91 | 0.8 | G95_50 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.79 | p138 | + | 3.828 | 186 (3.864-3.786) | 0.75 | 0.91 | 0.55 | G95_52 (1) | G90_38 (1) | G85_2 (39) | G80_1 (106) | + | |||
| 0.79 | p139 | + | 3.823 | 186 (3.864-3.786) | 0.8 | 0.78 | 0.45 | G95_53 (1) | G90_39 (1) | G85_2 (39) | G80_1 (106) | + | |||
| 0.74 | p154 | + | 3.76 | 188 (3.786-3.708) | 0.3 | 0.96 | 0.65 | G95_61 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.77 | p156 | + | 3.75 | 188 (3.786-3.708) | 0.68 | 0.7 | 0.35 | G95_62 (1) | G90_47 (1) | G85_32 (0) | G80_21 (0) | + | |||
| 0.86 | p160 | + | 3.718 | 188 (3.786-3.708) | 0.95 | 0.93 | 0.85 | G95_63 (1) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.91 | p162 | + | 3.739 | 189 (3.747-3.669) | 0.9 | 0.86 | 0.75 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.9 | p163 | + | 3.73 | 189 (3.747-3.669) | 0.95 | 0.89 | 0.85 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.9 | p165 | + | 3.709 | 189 (3.747-3.669) | 0.87 | 0.69 | 0.8 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.91 | p166 | + | 3.701 | 189 (3.747-3.669) | 0.82 | 0.61 | 0.7 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.8 | p208 | + | 3.547 | 194 (3.551-3.473) | 0.95 | 0.95 | 0.9 | G95_75 (0) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.93 | p209 | + | 3.54 | 194 (3.551-3.473) | 0.95 | 0.96 | 0.75 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.89 | p210 | + | 3.531 | 194 (3.551-3.473) | 0.96 | 0.93 | 0.8 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.9 | p211 | + | 3.524 | 194 (3.551-3.473) | 0.92 | 0.9 | 0.7 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.93 | p213 | + | 3.506 | 194 (3.551-3.473) | 0.97 | 0.96 | 0.75 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.92 | p214 | + | 3.491 | 194 (3.551-3.473) | 0.98 | 0.99 | 0.9 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.86 | p219 | + | 3.475 | 195 (3.512-3.434) | 0.5 | 0.96 | 0.85 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.88 | p221 | + | 3.464 | 195 (3.512-3.434) | 0.58 | 0.95 | 0.5 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.9 | p223 | + | 3.454 | 195 (3.512-3.434) | 0.81 | 0.98 | 0.45 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.83 | p234 | + | 3.396 | 197 (3.434-3.356) | 0.56 | 0.94 | 0.35 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.92 | p237 | + | 3.387 | 198 (3.395-3.317) | 0.96 | 0.96 | 0.7 | G95_2 (22) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.87 | p256 | + | 3.234 | 201 (3.277-3.199) | 0.94 | 0.95 | 0.55 | G95_83 (0) | G90_1 (36) | G85_2 (39) | G80_1 (106) | + | |||
| 0.72 | p261 | + | 3.231 | 202 (3.238-3.16) | 0.93 | 0.95 | 0.9 | G95_85 (0) | G90_61 (0) | G85_39 (0) | G80_1 (106) | + | |||
| 0.73 | p376 | + | 1.336 | 250 (1.361-1.283) | 0.98 | 0.89 | 0.95 | G95_7 (5) | G90_6 (6) | G85_4 (6) | G80_1 (106) | + | |||
| 0.73 | p377 | + | 1.325 | 250 (1.361-1.283) | 0.98 | 0.82 | 0.95 | G95_7 (5) | G90_6 (6) | G85_4 (6) | G80_1 (106) | + |
IDENTIFICATION ANALYSIS
| METABOLITE | TYPE | IC:MATCHING | IC:TOTAL | EC:MATCHING | EC:TOTAL | OTHERS | X | Y | Z | SCORE | FIG | |||
| Carnitine-7.0 (c:2/p:1) | M | 1 | 5 | 2 | 4 | 0 | -0.25 | 0.09 | 1.0 | 0.28 | + | |||
| Carnitine-7.0 (c:2/p:2) | M | 1 | 5 | 2 | 4 | 0 | -0.32 | 0.09 | 1.0 | 0.26 | + | |||
| glucose-D-1Phosphate-7.0 (c:4/p:1) | M | 1 | 3 | 4 | 5 | 0 | 0.46 | 0.85 | 1.0 | 0.77 | + | |||
| glucose-D-1Phosphate-7.0 (c:4/p:2) | M | 1 | 3 | 4 | 5 | 0 | 0.31 | 0.85 | 1.0 | 0.72 | + | |||
| glucose-D-1Phosphate-7.0 (c:4/p:3) | M | 1 | 3 | 4 | 5 | 0 | 0.22 | 0.85 | 1.0 | 0.69 | + | |||
| glucose-7.0 (c:4/p:5) | M | 2 | 14 | 3 | 5 | 0 | -0.11 | 0.74 | 1.0 | 0.54 | + | |||
| glucose-7.0 (c:4/p:6) | M | 3 | 14 | 3 | 5 | 0 | -0.12 | 0.74 | 1.0 | 0.54 | + | |||
| glucose-7.0 (c:4/p:7) | M | 1 | 14 | 3 | 5 | 0 | 0.33 | 0.74 | 1.0 | 0.69 | + | |||
| glucose-7.0 (c:4/p:8) | M | 1 | 14 | 3 | 5 | 0 | -0.09 | 0.74 | 1.0 | 0.55 | + | |||
| glucose-7.0 (c:4/p:9) | M | 1 | 14 | 3 | 5 | 0 | -0.11 | 0.74 | 1.0 | 0.54 | + |