PEAK 251 (3.261 ppm) DETAILS
GENERAL INFORMATION
PEAK DETECTION
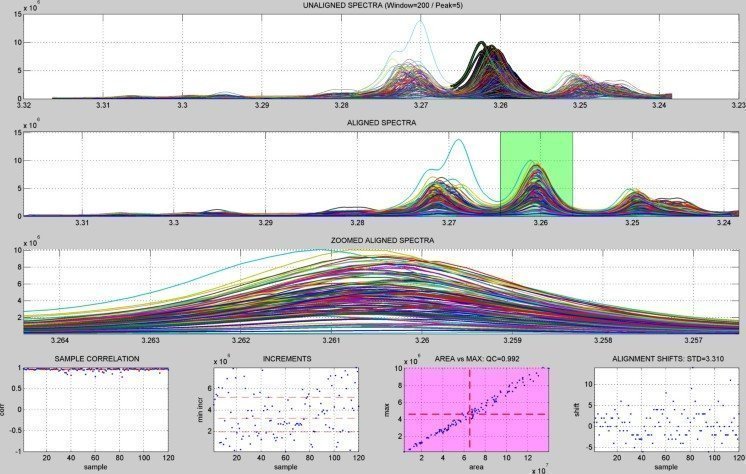
WINDOW ALIGNMENT

INTENSITY DISTRIBUTION

CORRELATION ANALYSIS

| CORRCOEF | ID_PEAK | INFO | PPM | ID_WINDOW | QC_SC | QC_IvsA | QC_IP | G_95 | G_90 | G_85 | G_80 | INFO | |||
| 0.7 | p9 | + | 8.346 | 70 (8.401-8.323) | 0.98 | 0.97 | 0.85 | G95_5 (7) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.73 | p18 | + | 7.871 | 82 (7.932-7.854) | 0.97 | 0.97 | 0.35 | G95_6 (5) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.72 | p19 | + | 7.884 | 83 (7.893-7.815) | 0.97 | 0.95 | 0.4 | G95_6 (5) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.72 | p46 | + | 6.11 | 128 (6.133-6.055) | 0.99 | 0.98 | 0.65 | G95_5 (7) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.73 | p47 | + | 6.101 | 128 (6.133-6.055) | 0.99 | 0.97 | 0.65 | G95_5 (7) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.73 | p48 | + | 5.923 | 132 (5.976-5.898) | 0.97 | 0.97 | 0.25 | G95_6 (5) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.74 | p49 | + | 5.916 | 132 (5.976-5.898) | 0.94 | 0.94 | 0.15 | G95_6 (5) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.74 | p53 | + | 5.908 | 133 (5.937-5.859) | 0.93 | 0.95 | 0.2 | G95_6 (5) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.72 | p55 | + | 5.894 | 134 (5.898-5.82) | 0.95 | 0.94 | 0.2 | G95_6 (5) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.74 | p77 | + | 4.449 | 170 (4.49-4.412) | 0.97 | 0.83 | 0.5 | G95_5 (7) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.73 | p78 | + | 4.441 | 170 (4.49-4.412) | 0.95 | 0.93 | 0.6 | G95_5 (7) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.77 | p83 | + | 4.293 | 174 (4.333-4.255) | 0.85 | 0.62 | 0.2 | G95_37 (1) | G90_30 (1) | G85_1 (69) | G80_1 (106) | + | |||
| 0.73 | p84 | + | 4.286 | 174 (4.333-4.255) | 0.9 | 0.74 | 0.4 | G95_5 (7) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.73 | p88 | + | 4.279 | 175 (4.294-4.216) | 0.89 | 0.69 | 0.35 | G95_38 (1) | G90_4 (16) | G85_1 (69) | G80_1 (106) | + | |||
| 0.97 | p227 | + | 3.437 | 196 (3.473-3.395) | 0.86 | 0.97 | 0.9 | G95_8 (4) | G90_8 (4) | G85_6 (4) | G80_2 (44) | + | |||
| 0.98 | p228 | + | 3.427 | 196 (3.473-3.395) | 0.95 | 0.95 | 0.95 | G95_8 (4) | G90_8 (4) | G85_6 (4) | G80_2 (44) | + | |||
| 0.94 | p229 | + | 3.416 | 196 (3.473-3.395) | 0.75 | 0.77 | 0.95 | G95_8 (4) | G90_8 (4) | G85_6 (4) | G80_2 (44) | + | |||
| 0.85 | p250 | + | 3.272 | 200 (3.316-3.238) | 0.87 | 0.85 | 0.95 | G95_82 (0) | G90_59 (0) | G85_6 (4) | G80_2 (44) | + | |||
| 0.96 | p252 | + | 3.25 | 200 (3.316-3.238) | 0.7 | 0.84 | 0.95 | G95_8 (4) | G90_8 (4) | G85_6 (4) | G80_2 (44) | + |
IDENTIFICATION ANALYSIS
| METABOLITE | TYPE | IC:MATCHING | IC:TOTAL | EC:MATCHING | EC:TOTAL | OTHERS | X | Y | Z | SCORE | FIG | |||
| Carnitine-7.0 (c:3/p:1) | M | 1 | 1 | 2 | 4 | 0 | 0.52 | 0.93 | 1.0 | 0.81 | + | |||
| glucose-7.0 (c:5/p:1) | M | 2 | 3 | 2 | 5 | 0 | 0.59 | 0.4 | 1.0 | 0.66 | + | |||
| glucose-7.0 (c:5/p:2) | M | 1 | 3 | 2 | 5 | 0 | 0.43 | 0.4 | 1.0 | 0.61 | + | |||
| glucose-7.0 (c:5/p:3) | M | 1 | 3 | 2 | 5 | 0 | 0.17 | 0.4 | 1.0 | 0.52 | + | |||
| histidine-7.0 (c:4/p:5) | M | 2 | 8 | 1 | 4 | 0 | 0.53 | 0.2 | 1.0 | 0.58 | + | |||
| histidine-7.0 (c:4/p:6) | M | 2 | 8 | 1 | 4 | 0 | 0.29 | 0.2 | 1.0 | 0.5 | + | |||
| phenylalanine-7.0 (c:3/p:3) | M | 2 | 4 | 1 | 4 | 0 | 0.62 | 0.17 | 1.0 | 0.6 | + | |||
| phenylalanine-7.0 (c:3/p:4) | M | 1 | 4 | 1 | 4 | 0 | 0.39 | 0.17 | 1.0 | 0.52 | ||||
| taurine-7.0 (c:2/p:1) | M | 1 | 4 | 2 | 2 | 0 | -0.55 | 1.0 | 1.0 | 0.48 | + | |||
| taurine-7.0 (c:2/p:2) | M | 2 | 4 | 2 | 2 | 0 | 0.62 | 1.0 | 1.0 | 0.87 | + | |||
| taurine-7.0 (c:2/p:3) | M | 2 | 4 | 2 | 2 | 0 | 0.79 | 1.0 | 1.0 | 0.93 | + | |||
| taurine-7.0 (c:2/p:4) | M | 1 | 4 | 2 | 2 | 0 | 0.06 | 1.0 | 1.0 | 0.69 | + |